Main | Research | Publications | Resources and miscellanea
Main Research Themes
This page is no longer maintained. Please visit: https://research.uni-leipzig.de/physics-of-evolution/
Understanding Phototrophic Growth
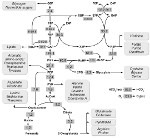 Our key challenge is to understand phototrophic growth of cyanobacteria in complex environments using methods from systems biology and theoretical biology.
We are interested in the organization and functioning of cyanobacterial metabolism.
Key research questions include the stoichiometric reconstruction of cyanobacterial metabolism, its comparison across different strains, the coordination of cyanobacterial metabolism in dynamic environments, in particular diurnal cycles, as well as resource allocation problems in dynamic metabolic networks.
We are interested in fast growth of cyanobacteria and investigate the limits of phototrophic growth using computational methods.
Our key challenge is to understand phototrophic growth of cyanobacteria in complex environments using methods from systems biology and theoretical biology.
We are interested in the organization and functioning of cyanobacterial metabolism.
Key research questions include the stoichiometric reconstruction of cyanobacterial metabolism, its comparison across different strains, the coordination of cyanobacterial metabolism in dynamic environments, in particular diurnal cycles, as well as resource allocation problems in dynamic metabolic networks.
We are interested in fast growth of cyanobacteria and investigate the limits of phototrophic growth using computational methods.
Further reading:
- Westermark S and Steuer R (2016) Toward Multiscale Models of Cyanobacterial Growth: A Modular Approach. Front. Bioeng. Biotechnol. 4:95. doi: 10.3389/fbioe.2016.00095
- Reimers AM, Knoop H, Bockmayr A, Steuer R. (2017) Cellular trade-offs and optimal resource allocation during cyanobacterial diurnal growth. Proc Natl Acad Sci U S A. pii: 201617508. doi: 10.1073/pnas.1617508114.
- H. Knoop, M. Gruendel, Y. Zilliges, R. Lehmann, S. Hoffmann, W. Lockau, R. Steuer. (2013) Flux Balance Analysis of Cyanobacterial Metabolism: The metabolic network of Synechocystis sp. PCC 6803. PLoS Comput Biol 9(6): e1003081. doi:10.1371/journal.pcbi.1003081
Cyanobacterial Biotechnology
 Cyanobacteria have attracted growing attention as potential host organisms for the production of valuable organic products. We develop computational methods to facilitate and enhance production of renewable bulk products using cyanobacteria. The aim is to integrate photosynthetic solar energy conversion and product formation, including engine-ready fuels, in a single biological process.
Past target products are ethanol (in collaboration with several academic and industrial partners, including Algenol Deutschland GmbH), as well as short chain (propane) and medium chain alkanes.
High-quality reconstructions of cyanobacterial metabolism are used to guide and support experimental efforts to increase and sustain product yield in cyanobacteria. The group participated in launching a start-up company to commercialize cultivation of cyanobacteria and microalgae at ultra-high densities (www.celldeg.com).
Cyanobacteria have attracted growing attention as potential host organisms for the production of valuable organic products. We develop computational methods to facilitate and enhance production of renewable bulk products using cyanobacteria. The aim is to integrate photosynthetic solar energy conversion and product formation, including engine-ready fuels, in a single biological process.
Past target products are ethanol (in collaboration with several academic and industrial partners, including Algenol Deutschland GmbH), as well as short chain (propane) and medium chain alkanes.
High-quality reconstructions of cyanobacterial metabolism are used to guide and support experimental efforts to increase and sustain product yield in cyanobacteria. The group participated in launching a start-up company to commercialize cultivation of cyanobacteria and microalgae at ultra-high densities (www.celldeg.com).
Further reading:
- Zavřel T, Červený J, Knoop H, Steuer R (2016) Optimizing cyanobacterial product synthesis: Meeting the challenges. Bioengineered 15:1-7.
- Knoop H, Steuer R. (2015) A computational analysis of stoichiometric constraints and trade-offs in cyanobacterial biofuel production. Front Bioeng Biotechnol. 3:47. doi: 10.3389/fbioe.2015.00047.
- Erdrich P, Knoop H, Steuer R, Klamt S. (2014) Cyanobacterial biofuels: new insights and strain design strategies revealed by computational modeling. Microb Cell Fact. 13(1):128.
The Nonlinear Dynamics of Metabolism
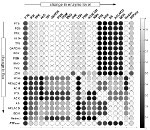 One of the most challenging goals of computational systems biology is the development of large-scale kinetic models of cellular pathways. However, for most cellular networks, detailed kinetic modeling is not possible due to lack of knowledge kinetic parameters. To overcome some of these problems, we are interested in novel methods that allow the elucidation of large-scale metabolic networks in the face of uncertain and incomplete information. Recent work includes novel approaches that provide a bridge between stoichiometric analysis and explicit kinetic simulations. Without requiring knowledge about the explicit functional form of the kinetic rate equations and parameters, these methods seek to describe the possible dynamics of cellular networks.
One of the most challenging goals of computational systems biology is the development of large-scale kinetic models of cellular pathways. However, for most cellular networks, detailed kinetic modeling is not possible due to lack of knowledge kinetic parameters. To overcome some of these problems, we are interested in novel methods that allow the elucidation of large-scale metabolic networks in the face of uncertain and incomplete information. Recent work includes novel approaches that provide a bridge between stoichiometric analysis and explicit kinetic simulations. Without requiring knowledge about the explicit functional form of the kinetic rate equations and parameters, these methods seek to describe the possible dynamics of cellular networks.
Further reading:
- Murabito E, Verma M, Bekker M, Bellomo D, Westerhoff HV, Teusink B, Steuer R. (2014 Monte-Carlo modeling of the central carbon metabolism of Lactococcus lactis: insights into metabolic regulation. PLoS One. 9(9):e106453.
- R. Steuer and B. H. Junker (2009) Computational Models of Metabolism: Stability and Regulation in Metabolic Networks. Advances in Chemical Physics, Volume 142. Rice, Stuart A. (editor), ISBN-10: 0-470-46499-2 ISBN-13: 978-0-470-46499-1 - John Wiley & Sons (2009)
- Steuer R, Gross T, Selbig J, Blasius B. (2006) Structural kinetic modeling of metabolic networks Proc Natl Acad Sci U S A. 103(32):11868-73.
Robustness and Principles of Cellular Signal Transduction
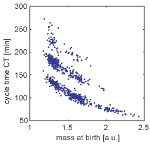 Noise and fluctuations are ubiquitous in living systems. Still, the interaction between complex biochemical regulatory networks and the inherent fluctuations ('noise') is only poorly understood. To elucidate the interrelation between noise and function, we investigate the implications of stochastic fluctuations on cellular regulatory systems, such as signal transduction networks, circadian clocks, or cell cycle control systems. Previous work includes the investigation of the effects of noise on a model of the eukaryotic cell cycle. The stochastic description leads to qualitative changes in the dynamic behavior, such as the emergence of noise-induced oscillations. Current research is focussed on a better understanding of robust and reliable information processing in living cells.
Noise and fluctuations are ubiquitous in living systems. Still, the interaction between complex biochemical regulatory networks and the inherent fluctuations ('noise') is only poorly understood. To elucidate the interrelation between noise and function, we investigate the implications of stochastic fluctuations on cellular regulatory systems, such as signal transduction networks, circadian clocks, or cell cycle control systems. Previous work includes the investigation of the effects of noise on a model of the eukaryotic cell cycle. The stochastic description leads to qualitative changes in the dynamic behavior, such as the emergence of noise-induced oscillations. Current research is focussed on a better understanding of robust and reliable information processing in living cells.
Further reading:
- Steuer R, Waldherr S, Sourjik V, Kollmann M (2011) Robust Signal Processing in Living Cells. PLoS Comput Biol 7(11): e1002218. doi:10.1371/journal.pcbi.1002218
- Ralf Steuer (2004) Effects of stochasticity in models of the cell cycle: From quantized cycle times to noise-induced oscillations J Theor Biol. 228(3):293-301.
Large-Scale Data Analysis and Interpretation of Metabolomics Data
 Metabolomic measurements provide a wealth of information about the biochemical status of cells, tissues and organs and play an important role to elucidate the function of novel genes. A remarkable inherent feature of cellular metabolism is that the concentrations of a small but significant number of metabolites are strongly correlated when measurements of biological replicates are performed. Drawing upon concepts of Nonlinear Dynamics and Computer Science, our work seeks to elucidate how comparative correlation analysis offers a way to exploit the intrinsic variability of metabolic networks to obtain significant additional information about the physiological state of the system.
Metabolomic measurements provide a wealth of information about the biochemical status of cells, tissues and organs and play an important role to elucidate the function of novel genes. A remarkable inherent feature of cellular metabolism is that the concentrations of a small but significant number of metabolites are strongly correlated when measurements of biological replicates are performed. Drawing upon concepts of Nonlinear Dynamics and Computer Science, our work seeks to elucidate how comparative correlation analysis offers a way to exploit the intrinsic variability of metabolic networks to obtain significant additional information about the physiological state of the system.
Further reading:
- Morgenthal K., Weckwerth W., Steuer R. (2006) Metabolomic networks in plants: Transitions from pattern recognition to biological interpretation. Biosystems. 83(2-3):108-17.
- R. Steuer, J. Kurths, O. Fiehn and W. Weckwerth (2003) Observing and interpreting correlations in metabolomic networks. Bioinformatics, Vol. 19 (8), 1019-1026